6886 results
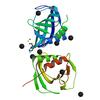
X-ray diffraction data for the Crystal structure of a thioesterase-like protein (CLOSPO_01618) from Clostridium sporogenes ATCC 15579 at 1.70 A resolution

First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.70 Å
R/Rfree: 0.18/0.20
Resolution: 1.70 Å
R/Rfree: 0.18/0.20
X-ray diffraction data for the Crystal structure of a DUF4909 family protein (SAV1798) from Staphylococcus aureus subsp. aureus Mu50 at 1.92 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.92 Å
R/Rfree: 0.20/0.24
Resolution: 1.92 Å
R/Rfree: 0.20/0.24
X-ray diffraction data for the Crystal structure of a protein with alpha-lytic protease prodomain-like fold (BDI_0842) from Parabacteroides distasonis ATCC 8503 at 1.40 A resolution


First author:
JOINT CENTER FOR STRUCTURAL GENOMICS (JCSG)
Resolution: 1.40 Å
R/Rfree: 0.13/0.17
Resolution: 1.40 Å
R/Rfree: 0.13/0.17
X-ray diffraction data for the Crystal structure of a TPR-domain containing protein (BDI_1685) from Parabacteroides distasonis ATCC 8503 at 2.26 A resolution


First author:
JOINT CENTER FOR STRUCTURAL GENOMICS (JCSG)
Resolution: 2.26 Å
R/Rfree: 0.18/0.21
Resolution: 2.26 Å
R/Rfree: 0.18/0.21
X-ray diffraction data for the Crystal structure of a putative esterase (BDI_1566) from Parabacteroides distasonis ATCC 8503 at 1.60 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.60 Å
R/Rfree: 0.16/0.19
Resolution: 1.60 Å
R/Rfree: 0.16/0.19
X-ray diffraction data for the Crystal structure of a putative cell adhesion protein (CLOSPO_03726) from Clostridium sporogenes ATCC 15579 at 1.95 A resolution


First author:
JOINT CENTER FOR STRUCTURAL GENOMICS (JCSG)
Resolution: 1.95 Å
R/Rfree: 0.17/0.21
Resolution: 1.95 Å
R/Rfree: 0.17/0.21
X-ray diffraction data for the Crystal structure of a conserved domain protein (SP_1775) from Streptococcus pneumoniae TIGR4 at 1.91 A resolution


First author:
JOINT CENTER FOR STRUCTURAL GENOMICS (JCSG)
Resolution: 1.91 Å
R/Rfree: 0.18/0.23
Resolution: 1.91 Å
R/Rfree: 0.18/0.23
X-ray diffraction data for the Crystal structure of a YbbR-like protein (SP_1560) from Streptococcus pneumoniae TIGR4 at 2.74 A resolution


First author:
JOINT CENTER FOR STRUCTURAL GENOMICS (JCSG)
Resolution: 2.74 Å
R/Rfree: 0.23/0.29
Resolution: 2.74 Å
R/Rfree: 0.23/0.29
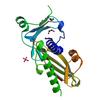
X-ray diffraction data for the Crystal structure of a DUF4467 family protein (SAV0303) from Staphylococcus aureus subsp. aureus Mu50 at 1.35 A resolution

First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.20 Å
R/Rfree: 0.14/0.17
Resolution: 1.20 Å
R/Rfree: 0.14/0.17
X-ray diffraction data for the Crystal structure of a cysteine-rich secretory protein (SAV1118) from Staphylococcus aureus subsp. aureus Mu50 at 1.90 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.90 Å
R/Rfree: 0.19/0.22
Resolution: 1.90 Å
R/Rfree: 0.19/0.22
X-ray diffraction data for the Crystal structure of a putative conserved lipoprotein (NT01CX_1156) from Clostridium novyi NT at 2.70 A resolution


First author:
JOINT CENTER FOR STRUCTURAL GENOMICS (JCSG)
Resolution: 2.70 Å
R/Rfree: 0.20/0.20
Resolution: 2.70 Å
R/Rfree: 0.20/0.20
X-ray diffraction data for the Crystal structure of a DUF1672 family protein (SAV1486) from Staphylococcus aureus subsp. aureus Mu50 at 1.80 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.80 Å
R/Rfree: 0.18/0.22
Resolution: 1.80 Å
R/Rfree: 0.18/0.22
X-ray diffraction data for the Crystal structure of a superantigen-like protein (SAV0433) from Staphylococcus aureus MU50 at 2.05 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.05 Å
R/Rfree: 0.19/0.24
Resolution: 2.05 Å
R/Rfree: 0.19/0.24
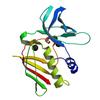
X-ray diffraction data for the Crystal structure of an exotoxin 6 (SAV0422) from Staphylococcus aureus subsp. aureus Mu50 at 2.25 A resolution

First author:
JOINT CENTER FOR STRUCTURAL GENOMICS (JCSG)
Resolution: 2.25 Å
R/Rfree: 0.19/0.24
Resolution: 2.25 Å
R/Rfree: 0.19/0.24
X-ray diffraction data for the Crystal structure of a putative peptide binding protein (RUMGNA_00914) from Ruminococcus gnavus ATCC 29149 at 1.60 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.60 Å
R/Rfree: 0.16/0.20
Resolution: 1.60 Å
R/Rfree: 0.16/0.20
X-ray diffraction data for the Crystal structure of a putative metal binding protein RUMGNA_00854 (ZP_02040092.1) from Ruminococcus gnavus ATCC 29149 at 1.30 A resolution


First author:
JOINT CENTER FOR STRUCTURAL GENOMICS (JCSG)
Resolution: 1.30 Å
R/Rfree: 0.14/0.19
Resolution: 1.30 Å
R/Rfree: 0.14/0.19
X-ray diffraction data for the Crystal structure of a DUF3887 family protein (RUMGNA_01855) from Ruminococcus gnavus ATCC 29149 at 2.25 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.25 Å
R/Rfree: 0.19/0.21
Resolution: 2.25 Å
R/Rfree: 0.19/0.21
X-ray diffraction data for the Crystal structure of a NTF2-like protein (EUBSIR_01394) from Eubacterium siraeum DSM 15702 at 2.56 A resolution


First author:
JOINT CENTER FOR STRUCTURAL GENOMICS (JCSG)
Resolution: 2.56 Å
R/Rfree: 0.25/0.30
Resolution: 2.56 Å
R/Rfree: 0.25/0.30
X-ray diffraction data for the Crystal structure of a protein with unknown function (CLOLEP_02462) from Clostridium leptum DSM 753 at 2.10 A resolution


First author:
JOINT CENTER FOR STRUCTURAL GENOMICS (JCSG)
Resolution: 2.10 Å
R/Rfree: 0.16/0.20
Resolution: 2.10 Å
R/Rfree: 0.16/0.20
X-ray diffraction data for the Crystal structure of a DUF5037 family protein (RUMGNA_01148) from Ruminococcus gnavus ATCC 29149 at 2.55 A resolution


First author:
Joint center for structural genomics (JCSG)
Resolution: 2.55 Å
R/Rfree: 0.19/0.22
Resolution: 2.55 Å
R/Rfree: 0.19/0.22
X-ray diffraction data for the Crystal structure of a putative transcription regulator (EUBSIR_01389) from Eubacterium siraeum DSM 15702 at 2.15 A resolution


First author:
JOINT CENTER FOR STRUCTURAL GENOMICS (JCSG)
Resolution: 2.15 Å
R/Rfree: 0.20/0.25
Resolution: 2.15 Å
R/Rfree: 0.20/0.25
X-ray diffraction data for the Crystal structure of a DHHW family protein (EUBSIR_00411) from Eubacterium siraeum DSM 15702 at 1.49 A resolution


First author:
JOINT CENTER FOR STRUCTURAL GENOMICS (JCSG)
Resolution: 1.49 Å
R/Rfree: 0.13/0.16
Resolution: 1.49 Å
R/Rfree: 0.13/0.16
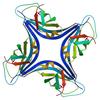
X-ray diffraction data for the Crystal structure of an apag protein (PA1934) from pseudomonas aeruginosa pao1 at 1.83 A resolution

First author:
Joint center for structural genomics (JCSG)
Resolution: 1.83 Å
R/Rfree: 0.19/0.22
Resolution: 1.83 Å
R/Rfree: 0.19/0.22
X-ray diffraction data for the Crystal structure of an apag protein (PA1934) from pseudomonas aeruginosa pao1 at 1.50 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.50 Å
R/Rfree: 0.18/0.22
Resolution: 1.50 Å
R/Rfree: 0.18/0.22
X-ray diffraction data for the Crystal structure of an immunoglobulin-like protein (PA1606) from PSEUDOMONAS AERUGINOSA at 1.50 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.50 Å
R/Rfree: 0.18/0.21
Resolution: 1.50 Å
R/Rfree: 0.18/0.21