6886 results
X-ray diffraction data for the Crystal structure of putative dioxygenase (YP_555069.1) from Burkholderia Xenovorans LB400 at 1.70 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.70 Å
R/Rfree: 0.18/0.22
Resolution: 1.70 Å
R/Rfree: 0.18/0.22
X-ray diffraction data for the Crystal structure of XisI protein-like (YP_323822.1) from Anabaena Variabilis ATCC 29413 at 1.85 A resolution


First author:
W.C. Hwang
Resolution: 1.85 Å
R/Rfree: 0.19/0.23
Resolution: 1.85 Å
R/Rfree: 0.19/0.23
X-ray diffraction data for the Crystal structure of fdxN element excision controlling factor XisI (YP_321976.1) from Anabaena Variabilis ATCC 29413 at 2.19 A resolution


First author:
W.C. Hwang
Resolution: 2.19 Å
R/Rfree: 0.20/0.24
Resolution: 2.19 Å
R/Rfree: 0.20/0.24
X-ray diffraction data for the Crystal structure of a duf1831 family protein (lp2179) from lactobacillus plantarum at 1.20 A resolution

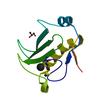
First author:
C. Bakolitsa
Resolution: 1.20 Å
R/Rfree: 0.12/0.15
Resolution: 1.20 Å
R/Rfree: 0.12/0.15
X-ray diffraction data for the Crystal structure of uncharacterized protein (YP_323524.1) from Anabaena variabilis ATCC 29413 at 2.11 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.11 Å
R/Rfree: 0.19/0.22
Resolution: 2.11 Å
R/Rfree: 0.19/0.22
X-ray diffraction data for the Crystal structure of a protein with a cupin-like fold and unknown function (YP_400729.1) from Synechococcus SP. PCC 7942 (Elongatus) at 2.50 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.50 Å
R/Rfree: 0.21/0.23
Resolution: 2.50 Å
R/Rfree: 0.21/0.23
X-ray diffraction data for the Crystal structure of a protein with unknown function from DUF155 family (YP_292156.1) from Prochlorococcus sp. NATL2A at 1.80 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.80 Å
R/Rfree: 0.18/0.21
Resolution: 1.80 Å
R/Rfree: 0.18/0.21
X-ray diffraction data for the Crystal structure of a xisi-like protein (npun_ar114) from nostoc punctiforme pcc 73102 at 2.30 A resolution

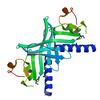
First author:
W.C. Hwang
Resolution: 2.30 Å
R/Rfree: 0.21/0.26
Resolution: 2.30 Å
R/Rfree: 0.21/0.26
X-ray diffraction data for the Crystal structure of an XisH family protein (ZP_00107633.1) from Nostoc punctiforme PCC 73102 at 1.60 A resolution


First author:
W.C. Hwang
Resolution: 1.60 Å
R/Rfree: 0.16/0.19
Resolution: 1.60 Å
R/Rfree: 0.16/0.19
X-ray diffraction data for the CRYSTAL STRUCTURE OF A XISI-LIKE PROTEIN (AVA_3825) FROM ANABAENA VARIABILIS ATCC 29413 AT 1.30 A RESOLUTION


First author:
W.C. Hwang
Resolution: 1.30 Å
R/Rfree: 0.18/0.20
Resolution: 1.30 Å
R/Rfree: 0.18/0.20
X-ray diffraction data for the CRYSTAL STRUCTURE OF A FDXN ELEMENT EXCISION CONTROLLING FACTOR PROTEIN (AVA_3312) FROM ANABAENA VARIABILIS AT 1.60 A RESOLUTION


First author:
W.C. Hwang
Resolution: 1.60 Å
R/Rfree: 0.14/0.16
Resolution: 1.60 Å
R/Rfree: 0.14/0.16
X-ray diffraction data for the Crystal structure of an uncharacterized conserved protein of COG5646 (ZP_00384875.1) from Lactobacillus casei ATCC 334 at 1.69 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.69 Å
R/Rfree: 0.17/0.20
Resolution: 1.69 Å
R/Rfree: 0.17/0.20
X-ray diffraction data for the Crystal structure of a protein from the duf1801 family (ydhg, bsu05750) from bacillus subtilis at 1.75 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.75 Å
R/Rfree: 0.17/0.22
Resolution: 1.75 Å
R/Rfree: 0.17/0.22
X-ray diffraction data for the Crystal structure of a protein of unknown function (NP_472245.1) from Listeria innocua at 1.65 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.65 Å
R/Rfree: 0.20/0.24
Resolution: 1.65 Å
R/Rfree: 0.20/0.24
X-ray diffraction data for the Crystal structure of an uncharacterized protein (yobk, bsu18990) from bacillus subtilis at 1.30 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.30 Å
R/Rfree: 0.14/0.18
Resolution: 1.30 Å
R/Rfree: 0.14/0.18
X-ray diffraction data for the CRYSTAL STRUCTURE OF A TETR-FAMILY TRANSCRIPTIONAL REGULATOR (ECA1819) FROM PECTOBACTERIUM ATROSEPTICUM AT 1.64 A RESOLUTION


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.64 Å
R/Rfree: 0.16/0.20
Resolution: 1.64 Å
R/Rfree: 0.16/0.20
X-ray diffraction data for the Crystal structure of predicted HD superfamily hydrolase involved in NAD metabolism (NP_347894.1) from Clostridium acetobutylicum at 1.50 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.50 Å
R/Rfree: 0.18/0.22
Resolution: 1.50 Å
R/Rfree: 0.18/0.22
X-ray diffraction data for the CRYSTAL STRUCTURE OF A PUTATIVE HD SUPERFAMILY HYDROLASE (BH1327) FROM BACILLUS HALODURANS AT 1.90 A RESOLUTION


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.90 Å
R/Rfree: 0.17/0.22
Resolution: 1.90 Å
R/Rfree: 0.17/0.22
X-ray diffraction data for the Crystal structure of Putative transcriptional regulator (JANN_22DEC04_CONTIG27_REVISED_GENE3569) from Jannaschia sp. CCS1 at 2.81 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.81 Å
R/Rfree: 0.20/0.25
Resolution: 2.81 Å
R/Rfree: 0.20/0.25
X-ray diffraction data for the Crystal structure of a tellurite resistance protein of cog3793 (npun_f6341) from nostoc punctiforme pcc 73102 at 1.85 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.85 Å
R/Rfree: 0.16/0.21
Resolution: 1.85 Å
R/Rfree: 0.16/0.21
X-ray diffraction data for the Crystal structure of ethanolamine ammonia-lyase heavy chain (YP_013784.1) from Listeria monocytogenes 4b F2365 at 2.15 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.15 Å
R/Rfree: 0.23/0.28
Resolution: 2.15 Å
R/Rfree: 0.23/0.28
X-ray diffraction data for the CRYSTAL STRUCTURE OF a PrpF family methylaconitate isomerase (PA0793) FROM PSEUDOMONAS AERUGINOSA AT 1.95 A RESOLUTION


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.95 Å
R/Rfree: 0.17/0.20
Resolution: 1.95 Å
R/Rfree: 0.17/0.20
X-ray diffraction data for the CRYSTAL STRUCTURE OF a protein belonging to the UPF0052 (SE_0549) FROM STAPHYLOCOCCUS EPIDERMIDIS ATCC 12228 AT 2.00 A RESOLUTION


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.00 Å
R/Rfree: 0.17/0.21
Resolution: 2.00 Å
R/Rfree: 0.17/0.21
X-ray diffraction data for the Crystal structure of a putative phospho transferase (sp_1565) from streptococcus pneumoniae tigr4 at 2.00 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.00 Å
R/Rfree: 0.20/0.24
Resolution: 2.00 Å
R/Rfree: 0.20/0.24
X-ray diffraction data for the Crystal structure of a protein member of the upf0052 family (bh3568) from bacillus halodurans at 2.60 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.60 Å
R/Rfree: 0.19/0.22
Resolution: 2.60 Å
R/Rfree: 0.19/0.22