1469 results
X-ray diffraction data for the Crystal structure of a biotin protein ligase-like protein of unknown function (tm1040_0394) from silicibacter sp. tm1040 at 1.70 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.70 Å
R/Rfree: 0.18/0.23
Resolution: 1.70 Å
R/Rfree: 0.18/0.23
X-ray diffraction data for the CRYSTAL STRUCTURE OF A PROTEIN WITH UNKNOWN FUNCTION FROM DUF3598 FAMILY (NPUN_R4044) FROM NOSTOC PUNCTIFORME PCC 73102 AT 1.75 A RESOLUTION


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.75 Å
R/Rfree: 0.17/0.21
Resolution: 1.75 Å
R/Rfree: 0.17/0.21
X-ray diffraction data for the Crystal structure of a binding protein component of ABC phosphonate transporter (PA3383) from Pseudomonas aeruginosa at 1.97 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.97 Å
R/Rfree: 0.16/0.19
Resolution: 1.97 Å
R/Rfree: 0.16/0.19
X-ray diffraction data for the CRYSTAL STRUCTURE OF A PUTATIVE THIOESTERASE (SSO2295) FROM SULFOLOBUS SOLFATARICUS AT 1.91 A RESOLUTION


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.91 Å
R/Rfree: 0.22/0.26
Resolution: 1.91 Å
R/Rfree: 0.22/0.26
X-ray diffraction data for the Crystal structure of an osmc-like hydroperoxide resistance protein (jann_2040) from jannaschia sp. ccs1 at 1.70 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.70 Å
R/Rfree: 0.14/0.16
Resolution: 1.70 Å
R/Rfree: 0.14/0.16
X-ray diffraction data for the Crystal structure of Fumarase of FUM-1 (NP_069927.1) from Archaeoglobus Fulgidus at 1.66 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.66 Å
R/Rfree: 0.17/0.20
Resolution: 1.66 Å
R/Rfree: 0.17/0.20
X-ray diffraction data for the CRYSTAL STRUCTURE OF A PROTEIN WITH UNKNOWN FUNCTION FROM DUF3224 FAMILY (SO_1590) FROM SHEWANELLA ONEIDENSIS MR-1 AT 1.84 A RESOLUTION

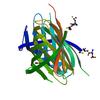
First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.84 Å
R/Rfree: 0.18/0.21
Resolution: 1.84 Å
R/Rfree: 0.18/0.21
X-ray diffraction data for the CRYSTAL STRUCTURE OF A PUTATIVE ANTIBIOTIC BIOSYNTHESIS MONOOXYGENASE (CC_2132) FROM CAULOBACTER CRESCENTUS CB15 AT 1.35 A RESOLUTION


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.35 Å
R/Rfree: 0.19/0.21
Resolution: 1.35 Å
R/Rfree: 0.19/0.21
X-ray diffraction data for the Crystal structure of carbamoylphosphate synthase large subunit (split gene in MJ) (ZP_00538348.1) from Exiguobacterium sp. 255-15 at 2.00 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.00 Å
R/Rfree: 0.19/0.24
Resolution: 2.00 Å
R/Rfree: 0.19/0.24
X-ray diffraction data for the Crystal structure of a two domain protein with unknown function (bt_3535) from bacteroides thetaiotaomicron vpi-5482 at 2.60 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.60 Å
R/Rfree: 0.20/0.24
Resolution: 2.60 Å
R/Rfree: 0.20/0.24
X-ray diffraction data for the Crystal structure of Putative SnoaL-like polyketide cyclase (YP_563807.1) from SHEWANELLA DENITRIFICANS OS-217 at 1.70 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.70 Å
R/Rfree: 0.19/0.24
Resolution: 1.70 Å
R/Rfree: 0.19/0.24
X-ray diffraction data for the Crystal structure of protein of unknown function with cystatin-like fold (NP_639274.1) from Xanthomonas campestris at 1.90 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.90 Å
R/Rfree: 0.17/0.20
Resolution: 1.90 Å
R/Rfree: 0.17/0.20
X-ray diffraction data for the Crystal structure of a putative polyketide cyclase (lferr_0659) from acidithiobacillus ferrooxidans atcc at 1.76 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.76 Å
R/Rfree: 0.17/0.21
Resolution: 1.76 Å
R/Rfree: 0.17/0.21
X-ray diffraction data for the Crystal structure of a pilus assembly protein CpaE (CC_2943) from Caulobacter crescentus CB15 at 1.75 A resolution (PSI Community Target, Shapiro)


First author:
JOINT CENTER FOR STRUCTURAL GENOMICS (JCSG)
Resolution: 1.75 Å
R/Rfree: 0.18/0.21
Resolution: 1.75 Å
R/Rfree: 0.18/0.21
X-ray diffraction data for the Crystal structure of a nitroreductase-like family protein (pnba, bh06130) from bartonella henselae str. houston-1 at 1.45 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.45 Å
R/Rfree: 0.15/0.17
Resolution: 1.45 Å
R/Rfree: 0.15/0.17
X-ray diffraction data for the Crystal structure of a Putative Pii-Like Signaling Protein (YP_323533.1) from ANABAENA VARIABILIS ATCC 29413 at 2.35 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.35 Å
R/Rfree: 0.22/0.27
Resolution: 2.35 Å
R/Rfree: 0.22/0.27
X-ray diffraction data for the Crystal structure of a SusD-like protein (BF3747) from Bacteroides fragilis NCTC 9343 at 2.70 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.70 Å
R/Rfree: 0.18/0.21
Resolution: 2.70 Å
R/Rfree: 0.18/0.21
X-ray diffraction data for the Crystal structure of a CheY-like protein (tadZ) from Pseudomonas aeruginosa PAO1 at 2.70 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.40 Å
R/Rfree: 0.20/0.23
Resolution: 2.40 Å
R/Rfree: 0.20/0.23
X-ray diffraction data for the CRYSTAL STRUCTURE OF A PUTATIVE ANTIBIOTIC BIOSYNTHESIS MONOOXYGENASE (SPO2313) FROM SILICIBACTER POMEROYI DSS-3 AT 1.30 A RESOLUTION


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.30 Å
R/Rfree: 0.13/0.15
Resolution: 1.30 Å
R/Rfree: 0.13/0.15
X-ray diffraction data for the Crystal structure of protein of unknown function with ferritin-like fold (YP_832262.1) from Arthrobacter sp. FB24 at 2.33 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.33 Å
R/Rfree: 0.21/0.26
Resolution: 2.33 Å
R/Rfree: 0.21/0.26
X-ray diffraction data for the Crystal structure of an alpha/beta hydrolase (YP_496220.1) from Novosphingobium aromaticivorans DSM 12444 at 1.50 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.50 Å
R/Rfree: 0.16/0.20
Resolution: 1.50 Å
R/Rfree: 0.16/0.20
X-ray diffraction data for the Crystal structure of a duf574 family protein (sav_2177) from streptomyces avermitilis ma-4680 at 1.80 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.80 Å
R/Rfree: 0.20/0.25
Resolution: 1.80 Å
R/Rfree: 0.20/0.25
X-ray diffraction data for the Crystal structure of 30S ribosomal protein S6 (TM0603) from Thermotoga maritima at 1.70 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.70 Å
R/Rfree: 0.18/0.24
Resolution: 1.70 Å
R/Rfree: 0.18/0.24
X-ray diffraction data for the Crystal structure of a putative two-domain sugar hydrolase (BACCAC_02064) from Bacteroides caccae ATCC 43185 at 1.35 A resolution


First author:
JOINT CENTER FOR STRUCTURAL GENOMICS (JCSG)
Resolution: 1.35 Å
R/Rfree: 0.11/0.13
Resolution: 1.35 Å
R/Rfree: 0.11/0.13
X-ray diffraction data for the Crystal structure of a putative neuraminidase (BACCAC_01090) from Bacteroides caccae ATCC 43185 at 1.90 A resolution (PSI Community Target)


First author:
JOINT CENTER FOR STRUCTURAL GENOMICS (JCSG)
Resolution: 1.90 Å
R/Rfree: 0.14/0.17
Resolution: 1.90 Å
R/Rfree: 0.14/0.17
X-ray diffraction data for the Crystal structure of a glycosyl hydrolase (BACOVA_03624) from Bacteroides ovatus at 2.30 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.30 Å
R/Rfree: 0.18/0.22
Resolution: 2.30 Å
R/Rfree: 0.18/0.22
X-ray diffraction data for the Crystal structure of a putative adhesin (BACOVA_04077) from Bacteroides ovatus at 2.50 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.50 Å
R/Rfree: 0.21/0.24
Resolution: 2.50 Å
R/Rfree: 0.21/0.24
X-ray diffraction data for the Crystal structure of a TetR family transcriptional regulator (Caur_2714) from CHLOROFLEXUS AURANTIACUS J-10-FL at 2.56 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.56 Å
R/Rfree: 0.21/0.24
Resolution: 2.56 Å
R/Rfree: 0.21/0.24
X-ray diffraction data for the CRYSTAL STRUCTURE OF AN UNCHARACTERIZED PROTEIN WITH A CYSTATIN-LIKE FOLD (CC_2572) FROM CAULOBACTER VIBRIOIDES AT 1.40 A RESOLUTION


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.40 Å
R/Rfree: 0.14/0.15
Resolution: 1.40 Å
R/Rfree: 0.14/0.15
X-ray diffraction data for the Crystal structure of an uncharacterized protein (PARMER_03598) from Parabacteroides merdae ATCC 43184 at 1.50 A resolution


First author:
JOINT CENTER FOR STRUCTURAL GENOMICS (JCSG)
Resolution: 1.50 Å
R/Rfree: 0.17/0.19
Resolution: 1.50 Å
R/Rfree: 0.17/0.19
X-ray diffraction data for the Crystal structure of a putative cell surface protein (BACOVA_01565) from Bacteroides ovatus ATCC 8483 at 2.05 A resolution


First author:
JOINT CENTER FOR STRUCTURAL GENOMICS (JCSG)
Resolution: 2.05 Å
R/Rfree: 0.18/0.22
Resolution: 2.05 Å
R/Rfree: 0.18/0.22
X-ray diffraction data for the Crystal structure of a putative hydrolase (lpg1103) from Legionella pneumophila subsp. pneumophila str. Philadelphia 1 at 1.15 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.15 Å
R/Rfree: 0.14/0.16
Resolution: 1.15 Å
R/Rfree: 0.14/0.16
X-ray diffraction data for the Crystal structure of putative MarR-like transcription regulator (NP_978771.1) from Bacillus cereus ATCC 10987 at 2.38 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.38 Å
R/Rfree: 0.23/0.25
Resolution: 2.38 Å
R/Rfree: 0.23/0.25
X-ray diffraction data for the CRYSTAL STRUCTURE OF A PUTATIVE SUGAR PHOSPHATE ISOMERASE/EPIMERASE (AVA4194) FROM ANABAENA VARIABILIS ATCC 29413 AT 1.78 A RESOLUTION


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.78 Å
R/Rfree: 0.18/0.22
Resolution: 1.78 Å
R/Rfree: 0.18/0.22
X-ray diffraction data for the Crystal structure of a duf1048 protein with a left-handed superhelix fold (bce_3448) from bacillus cereus atcc 10987 at 2.05 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.05 Å
R/Rfree: 0.18/0.23
Resolution: 2.05 Å
R/Rfree: 0.18/0.23
X-ray diffraction data for the Crystal structure of SusD-like carbohydrate binding protein (YP_001298396.1) from Bacteroides vulgatus ATCC 8482 at 1.70 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.70 Å
R/Rfree: 0.15/0.17
Resolution: 1.70 Å
R/Rfree: 0.15/0.17
X-ray diffraction data for the CRYSTAL STRUCTURE OF A DIMERIC FERREDOXIN-LIKE PROTEIN (JCVI_PEP_1096672785533) FROM UNCULTURED MARINE ORGANISM AT 2.20 A RESOLUTION


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.20 Å
R/Rfree: 0.23/0.29
Resolution: 2.20 Å
R/Rfree: 0.23/0.29
X-ray diffraction data for the Crystal structure of a cupin-like protein (tm1010) from thermotoga maritima msb8 at 1.90 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.90 Å
R/Rfree: 0.16/0.22
Resolution: 1.90 Å
R/Rfree: 0.16/0.22
X-ray diffraction data for the Crystal structure of a putative periplasmic protein of unknown function (bvu_2443) from bacteroides vulgatus atcc 8482 at 1.64 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.64 Å
R/Rfree: 0.17/0.21
Resolution: 1.64 Å
R/Rfree: 0.17/0.21
X-ray diffraction data for the CRYSTAL STRUCTURE OF a putative oxidoreductase of the DsrE/DsrF-like family (SSO1126) FROM SULFOLOBUS SOLFATARICUS P2 AT 2.09 A RESOLUTION


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.09 Å
R/Rfree: 0.18/0.20
Resolution: 2.09 Å
R/Rfree: 0.18/0.20
X-ray diffraction data for the Crystal structure of a putative inositol catabolism protein iole (iole, lp_3607) from lactobacillus plantarum wcfs1 at 1.85 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.85 Å
R/Rfree: 0.17/0.21
Resolution: 1.85 Å
R/Rfree: 0.17/0.21
X-ray diffraction data for the CRYSTAL STRUCTURE OF A PUTATIVE ACETYLACETONE DIOXYGENASE (MPE_A3659) FROM METHYLIBIUM PETROLEIPHILUM PM1 AT 1.50 A RESOLUTION


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.50 Å
R/Rfree: 0.18/0.21
Resolution: 1.50 Å
R/Rfree: 0.18/0.21
X-ray diffraction data for the Crystal structure of a duf89 family protein (ph1575) from pyrococcus horikoshii at 2.04 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.04 Å
R/Rfree: 0.14/0.18
Resolution: 2.04 Å
R/Rfree: 0.14/0.18
X-ray diffraction data for the Crystal structure of Gamma-glutamyl phosphate reductase (yor323c) from Saccharomyces cerevisiae at 2.40 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.29 Å
R/Rfree: 0.21/0.25
Resolution: 2.29 Å
R/Rfree: 0.21/0.25
X-ray diffraction data for the Crystal structure of Putative sugar binding protein (YP_001300177.1) from Bacteroides vulgatus ATCC 8482 at 1.70 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.70 Å
R/Rfree: 0.15/0.19
Resolution: 1.70 Å
R/Rfree: 0.15/0.19
X-ray diffraction data for the Crystal structure of a putative 3-keto-5-aminohexanoate cleavage enzyme (reut_c6226) from ralstonia eutropha jmp134 at 1.72 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.72 Å
R/Rfree: 0.15/0.18
Resolution: 1.72 Å
R/Rfree: 0.15/0.18
X-ray diffraction data for the Crystal structure of a putative 4-methylmuconolactone methylisomerase (YP_295714.1) from Ralstonia eutropha JMP134 at 2.00 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.00 Å
R/Rfree: 0.17/0.20
Resolution: 2.00 Å
R/Rfree: 0.17/0.20
X-ray diffraction data for the Crystal structure of a putative dipeptidyl-peptidase VI (BT_1314) from Bacteroides thetaiotaomicron VPI-5482 at 1.75 A resolution


First author:
JOINT CENTER FOR STRUCTURAL GENOMICS (JCSG)
Resolution: 1.75 Å
R/Rfree: 0.15/0.18
Resolution: 1.75 Å
R/Rfree: 0.15/0.18
X-ray diffraction data for the CRYSTAL STRUCTURE OF A PUTATIVE DEHYDRATASE FROM THE NTF2-LIKE FAMILY (SAV_4671) FROM STREPTOMYCES AVERMITILIS AT 2.10 A RESOLUTION


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.10 Å
R/Rfree: 0.20/0.24
Resolution: 2.10 Å
R/Rfree: 0.20/0.24
X-ray diffraction data for the Crystal structure of a ribonucleotide reductase M2 B (RNRR2) from Homo sapiens at 2.20 A resolution


First author:
Partnership for T-Cell Biology (TCELL) Joint Center for Structural Genomics (JCSG)
Resolution: 2.20 Å
R/Rfree: 0.19/0.22
Resolution: 2.20 Å
R/Rfree: 0.19/0.22
X-ray diffraction data for the Crystal structure of acetoacetate decarboxylase (ADC) (YP_094708.1) from Legionella pneumophila subsp. pneumophila str. Philadelphia 1 at 1.60 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.60 Å
R/Rfree: 0.19/0.23
Resolution: 1.60 Å
R/Rfree: 0.19/0.23
X-ray diffraction data for the CRYSTAL STRUCTURE OF A PUTATIVE THIOESTERASE, PHENYLACETIC ACID DEGRADATION-RELATED PROTEIN (REUT_B4779) FROM RALSTONIA EUTROPHA JMP134 AT 2.20 A RESOLUTION


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.20 Å
R/Rfree: 0.18/0.22
Resolution: 2.20 Å
R/Rfree: 0.18/0.22
X-ray diffraction data for the Crystal structure of a putative monooxygenase (YP_001095275.1) from Shewanella loihica PV-4 at 1.26 A resolution

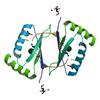
First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.26 Å
R/Rfree: 0.14/0.16
Resolution: 1.26 Å
R/Rfree: 0.14/0.16
X-ray diffraction data for the Crystal structure of a xanthine dehydrogenase (BH1974) from Bacillus halodurans at 2.80 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.80 Å
R/Rfree: 0.20/0.22
Resolution: 2.80 Å
R/Rfree: 0.20/0.22
X-ray diffraction data for the Crystal structure of a DJ-1 (PARK7) from Homo sapiens at 1.23 A resolution


First author:
Partnership for Nuclear Receptor Signaling Code Biology (NHRs) Joint Center for Structural Genomics (JCSG)
Resolution: 1.23 Å
R/Rfree: 0.12/0.13
Resolution: 1.23 Å
R/Rfree: 0.12/0.13
X-ray diffraction data for the Crystal structure of a DUF4465 family protein (BACCAC_02373) from Bacteroides caccae ATCC 43185 at 2.05 A resolution


First author:
JOINT CENTER FOR STRUCTURAL GENOMICS (JCSG)
Resolution: 2.05 Å
R/Rfree: 0.16/0.19
Resolution: 2.05 Å
R/Rfree: 0.16/0.19
X-ray diffraction data for the Crystal structure of a putative 2,3,4,5-tetrahydropyridine-2-carboxylate n-succinyltransferase (cj1605c, dapd) from campylobacter jejuni at 1.90 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.90 Å
R/Rfree: 0.16/0.19
Resolution: 1.90 Å
R/Rfree: 0.16/0.19
X-ray diffraction data for the Crystal structure of putative cystathionine beta-lyase involved in aluminum resistance (NP_470671.1) from LISTERIA INNOCUA at 2.12 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.12 Å
R/Rfree: 0.15/0.19
Resolution: 2.12 Å
R/Rfree: 0.15/0.19
X-ray diffraction data for the CRYSTAL STRUCTURE OF A UBIQUINONE/MENAQUINONE BIOSYNTHESIS METHYLTRANSFERASE-RELATED PROTEIN (TM1389) FROM THERMOTOGA MARITIMA MSB8 AT 2.35 A RESOLUTION


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.35 Å
R/Rfree: 0.17/0.20
Resolution: 2.35 Å
R/Rfree: 0.17/0.20
X-ray diffraction data for the Crystal structure of Homoserine O-succinyltransferase (EC 2.3.1.46) (Homoserine O-transsuccinylase) (HTS) (tm0881) from THERMOTOGA MARITIMA at 2.52 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.52 Å
R/Rfree: 0.19/0.27
Resolution: 2.52 Å
R/Rfree: 0.19/0.27
X-ray diffraction data for the Crystal structure of Peptide chain release factor 1 (RF-1) (SMU.1085) from Streptococcus mutans at 2.34 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.34 Å
R/Rfree: 0.22/0.28
Resolution: 2.34 Å
R/Rfree: 0.22/0.28
X-ray diffraction data for the Crystal structure of a lipoprotein, YaeC family (EF3198) from Enterococcus faecalis V583 at 1.80 A resolution


First author:
JOINT CENTER FOR STRUCTURAL GENOMICS (JCSG)
Resolution: 1.80 Å
R/Rfree: 0.18/0.21
Resolution: 1.80 Å
R/Rfree: 0.18/0.21
X-ray diffraction data for the Crystal structure of a n-acetylneuraminic acid phosphatase (nanp) from mus musculus at 1.90 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.90 Å
R/Rfree: 0.21/0.26
Resolution: 1.90 Å
R/Rfree: 0.21/0.26
X-ray diffraction data for the CRYSTAL STRUCTURE OF A TYW3 METHYLTRANSFERASE-LIKE PROTEIN (AF_2059) FROM ARCHAEOGLOBUS FULGIDUS DSM 4304 AT 1.95 A RESOLUTION


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.95 Å
R/Rfree: 0.20/0.25
Resolution: 1.95 Å
R/Rfree: 0.20/0.25
X-ray diffraction data for the CRYSTAL STRUCTURE OF A PUTATIVE FMN-BINDING PROTEIN (TA1372) FROM THERMOPLASMA ACIDOPHILUM AT 1.80 A RESOLUTION


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.80 Å
R/Rfree: 0.19/0.23
Resolution: 1.80 Å
R/Rfree: 0.19/0.23
X-ray diffraction data for the Crystal structure of a putative adenylate cyclase (bh2851) from bacillus halodurans at 2.12 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.12 Å
R/Rfree: 0.23/0.27
Resolution: 2.12 Å
R/Rfree: 0.23/0.27
X-ray diffraction data for the Crystal structure of a putative acylhydrolase (BF3764) from Bacteroides fragilis NCTC 9343 at 2.00 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.00 Å
R/Rfree: 0.19/0.21
Resolution: 2.00 Å
R/Rfree: 0.19/0.21
X-ray diffraction data for the Crystal structure of a DUF3299 family protein (PA4066) from Pseudomonas aeruginosa PAO1 at 2.45 A resolution

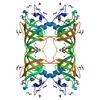
First author:
JOINT CENTER FOR STRUCTURAL GENOMICS (JCSG)
Resolution: 2.45 Å
R/Rfree: 0.17/0.21
Resolution: 2.45 Å
R/Rfree: 0.17/0.21
X-ray diffraction data for the Crystal structure of SAM-dependent O-methyltransferase (TM0748) from Thermotoga maritima at 1.65 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.65 Å
R/Rfree: 0.16/0.19
Resolution: 1.65 Å
R/Rfree: 0.16/0.19
X-ray diffraction data for the Crystal structure of Activation (AdoMet binding) domain of Methionine synthase (TM0269) from Thermotoga maritima at 2.2 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.30 Å
R/Rfree: 0.25/0.29
Resolution: 2.30 Å
R/Rfree: 0.25/0.29
X-ray diffraction data for the Crystal structure of an antioxidant defense protein (mlr4105) from mesorhizobium loti maff303099 at 2.00 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.00 Å
R/Rfree: 0.17/0.23
Resolution: 2.00 Å
R/Rfree: 0.17/0.23
X-ray diffraction data for the Crystal structure of cyclopropane-fatty-acyl-phospholipid synthase-like protein (YP_807781.1) from Lactobacillus casei ATCC 334 at 1.85 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.85 Å
R/Rfree: 0.19/0.24
Resolution: 1.85 Å
R/Rfree: 0.19/0.24
X-ray diffraction data for the CRYSTAL STRUCTURE OF A NTF-2 LIKE PROTEIN OF UNKNOWN FUNCTION (SO_0125) FROM SHEWANELLA ONEIDENSIS MR-1 AT 1.70 A RESOLUTION

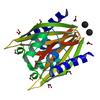
First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.70 Å
R/Rfree: 0.18/0.21
Resolution: 1.70 Å
R/Rfree: 0.18/0.21
X-ray diffraction data for the Crystal structure of a putative heme oxygenase (alr5027) from nostoc sp. pcc 7120 at 1.50 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.50 Å
R/Rfree: 0.15/0.18
Resolution: 1.50 Å
R/Rfree: 0.15/0.18
X-ray diffraction data for the Crystal structure of glutathione S-transferase (NP_416804.1) from Escherichia coli K12 at 1.85 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.85 Å
R/Rfree: 0.16/0.20
Resolution: 1.85 Å
R/Rfree: 0.16/0.20
X-ray diffraction data for the Crystal structure of an Imelysin-like protein (Psyc_1802) from PSYCHROBACTER ARCTICUM 273-4 at 2.15 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.15 Å
R/Rfree: 0.18/0.22
Resolution: 2.15 Å
R/Rfree: 0.18/0.22
X-ray diffraction data for the CRYSTAL STRUCTURE OF a PrpF family methylaconitate isomerase (PA0793) FROM PSEUDOMONAS AERUGINOSA AT 1.95 A RESOLUTION


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.95 Å
R/Rfree: 0.17/0.20
Resolution: 1.95 Å
R/Rfree: 0.17/0.20
X-ray diffraction data for the Crystal structure of a putative cell adhesion protein (BDI_3519) from Parabacteroides distasonis ATCC 8503 at 2.34 A resolution (PSI Community Target, Nakayama)


First author:
JOINT CENTER FOR STRUCTURAL GENOMICS (JCSG)
Resolution: 2.34 Å
R/Rfree: 0.18/0.23
Resolution: 2.34 Å
R/Rfree: 0.18/0.23
X-ray diffraction data for the CRYSTAL STRUCTURE OF a DUF72 family protein (EF0366) FROM ENTEROCOCCUS FAECALIS V583 AT 2.52 A RESOLUTION


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.52 Å
R/Rfree: 0.23/0.26
Resolution: 2.52 Å
R/Rfree: 0.23/0.26
X-ray diffraction data for the Crystal structure of a protein of unknown function with a cystatin-like fold (saro_2880) from novosphingobium aromaticivorans dsm at 1.30 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.30 Å
R/Rfree: 0.12/0.14
Resolution: 1.30 Å
R/Rfree: 0.12/0.14
X-ray diffraction data for the CRYSTAL STRUCTURE OF A PUTATIVE LIPID BINDING PROTEIN (GSU0061) FROM GEOBACTER SULFURREDUCENS PCA AT 1.80 A RESOLUTION


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.80 Å
R/Rfree: 0.19/0.22
Resolution: 1.80 Å
R/Rfree: 0.19/0.22
X-ray diffraction data for the CRYSTAL STRUCTURE OF A PUTATIVE TRNA-(MS(2)IO(6)A)-HYDROXYLASE (PP_2188) FROM PSEUDOMONAS PUTIDA KT2440 AT 2.05 A RESOLUTION


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.05 Å
R/Rfree: 0.17/0.23
Resolution: 2.05 Å
R/Rfree: 0.17/0.23
X-ray diffraction data for the Crystal structure of a DUF2874 family protein (BACOVA_02504) from Bacteroides ovatus ATCC 8483 at 1.62 A resolution


First author:
JOINT CENTER FOR STRUCTURAL GENOMICS (JCSG)
Resolution: 1.62 Å
R/Rfree: 0.17/0.20
Resolution: 1.62 Å
R/Rfree: 0.17/0.20
X-ray diffraction data for the Crystal structure of Predicted metal-dependent phosphoesterase (PHP family) (tm0559) from THERMOTOGA MARITIMA at 2.40 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.40 Å
R/Rfree: 0.17/0.22
Resolution: 2.40 Å
R/Rfree: 0.17/0.22
X-ray diffraction data for the Crystal structure of a Putative gamma-D-glutamyl-L-diamino acid endopeptidase (DVU_0896) from DESULFOVIBRIO VULGARIS HILDENBOROUGH at 1.75 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.75 Å
R/Rfree: 0.14/0.17
Resolution: 1.75 Å
R/Rfree: 0.14/0.17
X-ray diffraction data for the Crystal structure of Sugar phosphate isomerase/epimerase (YP_001303399.1) from Parabacteroides distasonis ATCC 8503 at 1.70 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.70 Å
R/Rfree: 0.15/0.17
Resolution: 1.70 Å
R/Rfree: 0.15/0.17
X-ray diffraction data for the Crystal structure of uncharacterized protein conserved in bacteria with a cystatin-like fold (YP_168589.1) from SILICIBACTER POMEROYI DSS-3 at 2.00 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.00 Å
R/Rfree: 0.18/0.19
Resolution: 2.00 Å
R/Rfree: 0.18/0.19
X-ray diffraction data for the Crystal structure of a DUF4886 family protein (BACUNI_01406) from Bacteroides uniformis ATCC 8492 at 1.50 A resolution


First author:
JOINT CENTER FOR STRUCTURAL GENOMICS (JCSG)
Resolution: 1.50 Å
R/Rfree: 0.15/0.18
Resolution: 1.50 Å
R/Rfree: 0.15/0.18
X-ray diffraction data for the Crystal structure of pyruvate oxidoreductase subunit PORC (EC 1.2.7.1) (TM0015) from Thermotoga maritima at 2.12 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.12 Å
R/Rfree: 0.22/0.25
Resolution: 2.12 Å
R/Rfree: 0.22/0.25
X-ray diffraction data for the Crystal structure of a peptidyl-prolyl cis-trans isomerase E (PPIE) from Homo sapiens at 2.50 A resolution


First author:
Partnership for T-Cell Biology JOINT CENTER FOR STRUCTURAL GENOMICS (JCSG)
Resolution: 2.50 Å
R/Rfree: 0.19/0.21
Resolution: 2.50 Å
R/Rfree: 0.19/0.21
X-ray diffraction data for the Crystal structure of a DUF4738 family protein (BACOVA_05496) from Bacteroides ovatus ATCC 8483 at 2.15 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.15 Å
R/Rfree: 0.21/0.23
Resolution: 2.15 Å
R/Rfree: 0.21/0.23
X-ray diffraction data for the CRYSTAL STRUCTURE OF A PUTATIVE THIOESTERASE II (TFU_2367) FROM THERMOBIFIDA FUSCA YX AT 2.45 A RESOLUTION


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.16 Å
R/Rfree: 0.18/0.24
Resolution: 2.16 Å
R/Rfree: 0.18/0.24
X-ray diffraction data for the Crystal structure of a duf1611 family protein (ava_3511) from anabaena variabilis atcc 29413 at 2.30 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.30 Å
R/Rfree: 0.18/0.21
Resolution: 2.30 Å
R/Rfree: 0.18/0.21
X-ray diffraction data for the Crystal structure of Putative TetR-family transcriptional regulator (YP_752756.1) from Syntrophomonas wolfei str. Goettingen at 2.07 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.07 Å
R/Rfree: 0.19/0.23
Resolution: 2.07 Å
R/Rfree: 0.19/0.23
X-ray diffraction data for the Crystal structure of a RNA-binding protein 39 (RBM39) in complex with fragment of splicing factor (U2AF) from Mus musculus at 2.20 A resolution


First author:
Partnership for T-Cell Biology (TCELL) Joint Center for Structural Genomics (JCSG)
Resolution: 2.20 Å
R/Rfree: 0.17/0.20
Resolution: 2.20 Å
R/Rfree: 0.17/0.20
X-ray diffraction data for the Crystal structure of an ethd-like protein (reut_b5694) from ralstonia eutropha jmp134 at 2.10 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.10 Å
R/Rfree: 0.18/0.21
Resolution: 2.10 Å
R/Rfree: 0.18/0.21
X-ray diffraction data for the Crystal structure of a vinyl-4-reductase family protein (mj_1460) from methanocaldococcus jannaschii dsm at 1.90 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.90 Å
R/Rfree: 0.19/0.25
Resolution: 1.90 Å
R/Rfree: 0.19/0.25
X-ray diffraction data for the Crystal structure of dimeric protein of unknown function and ferredoxin-like fold (YP_212648.1) from Bacteroides fragilis NCTC 9343 at 1.85 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.85 Å
R/Rfree: 0.17/0.21
Resolution: 1.85 Å
R/Rfree: 0.17/0.21
X-ray diffraction data for the Crystal structure of an atmB (putative membrane lipoprotein) from Streptococcus mutans UA159 at 1.91 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.91 Å
R/Rfree: 0.18/0.23
Resolution: 1.91 Å
R/Rfree: 0.18/0.23
X-ray diffraction data for the Crystal structure of a putative alpha-galactosidase (BF1418) from Bacteroides fragilis NCTC 9343 at 1.57 A resolution


First author:
JOINT CENTER FOR STRUCTURAL GENOMICS (JCSG)
Resolution: 1.57 Å
R/Rfree: 0.15/0.17
Resolution: 1.57 Å
R/Rfree: 0.15/0.17