1469 results
X-ray diffraction data for the Crystal structure of a putative transposase (dr_0177) from deinococcus radiodurans r1 at 1.90 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.90 Å
R/Rfree: 0.16/0.19
Resolution: 1.90 Å
R/Rfree: 0.16/0.19
X-ray diffraction data for the Crystal structure of Putative NADH dehydrogenase/NAD(P)H nitroreductase (NP_809094.1) from BACTEROIDES THETAIOTAOMICRON VPI-5482 at 1.54 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.54 Å
R/Rfree: 0.15/0.18
Resolution: 1.54 Å
R/Rfree: 0.15/0.18
X-ray diffraction data for the Crystal structure of Iron-containing alcohol dehydrogenase (NP_602249.1) from Corynebacterium glutamicum ATCC 13032 KITASATO at 2.07 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.07 Å
R/Rfree: 0.17/0.22
Resolution: 2.07 Å
R/Rfree: 0.17/0.22
X-ray diffraction data for the Crystal structure of a sam dependent methyl-transferase type 12 family protein (eca1738) from pectobacterium atrosepticum scri1043 at 1.85 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.85 Å
R/Rfree: 0.15/0.18
Resolution: 1.85 Å
R/Rfree: 0.15/0.18
X-ray diffraction data for the Crystal structure of Protein of unknown function (DUF861) with a RmlC-like cupin fold (17741406) from AGROBACTERIUM TUMEFACIENS str. C58 (Dupont) at 1.64 A resolution

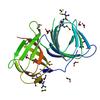
First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.64 Å
R/Rfree: 0.14/0.16
Resolution: 1.64 Å
R/Rfree: 0.14/0.16
X-ray diffraction data for the Crystal structure of a cupin domain (bp2299) from bordetella pertussis tohama i at 1.80 A resolution

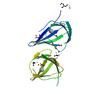
First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.80 Å
R/Rfree: 0.15/0.20
Resolution: 1.80 Å
R/Rfree: 0.15/0.20
X-ray diffraction data for the Crystal structure of a duf3244 family protein with an immunoglobulin-like beta-sandwich fold (bvu_0276) from bacteroides vulgatus atcc 8482 at 1.70 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.70 Å
R/Rfree: 0.17/0.22
Resolution: 1.70 Å
R/Rfree: 0.17/0.22
X-ray diffraction data for the Crystal structure of 3-oxoacyl-(acyl carrier protein) reductase (TM1169) from Thermotoga maritima at 2.50 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.50 Å
R/Rfree: 0.19/0.25
Resolution: 2.50 Å
R/Rfree: 0.19/0.25
X-ray diffraction data for the Crystal structure of Glia maturation factor-gamma (GMFG) from Mus musculus at 1.50 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.35 Å
R/Rfree: 0.16/0.19
Resolution: 1.35 Å
R/Rfree: 0.16/0.19
X-ray diffraction data for the CRYSTAL STRUCTURE OF a putativeTenA family transcriptional regulator (BT_3146) FROM BACTEROIDES THETAIOTAOMICRON VPI-5482 AT 1.88 A RESOLUTION


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.88 Å
R/Rfree: 0.14/0.17
Resolution: 1.88 Å
R/Rfree: 0.14/0.17
X-ray diffraction data for the Crystal structure of NADH:FMN oxidoreductase like protein in complex with FMN (YP_544701.1) from METHYLOBACILLUS FLAGELLATUS KT at 1.40 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.40 Å
R/Rfree: 0.12/0.15
Resolution: 1.40 Å
R/Rfree: 0.12/0.15
X-ray diffraction data for the Crystal structure of putative monooxygenase (YP_193413.1) from Lactobacillus acidophilus NCFM at 1.55 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.55 Å
R/Rfree: 0.18/0.20
Resolution: 1.55 Å
R/Rfree: 0.18/0.20
X-ray diffraction data for the Crystal structure of a Multidomain protein including DUF1735 (BACOVA_03322) from Bacteroides ovatus at 2.80 A resolution


First author:
Joint center for structural genomics (jcsg)
Resolution: 2.80 Å
R/Rfree: 0.20/0.23
Resolution: 2.80 Å
R/Rfree: 0.20/0.23
X-ray diffraction data for the Crystal structure of Putative antibiotic biosynthesis monooxygenase (NP_810307.1) from Bacteriodes thetaiotaomicron VPI-5482 at 1.30 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.30 Å
R/Rfree: 0.14/0.17
Resolution: 1.30 Å
R/Rfree: 0.14/0.17
X-ray diffraction data for the Crystal structure of a putative sulfatase (NP_810509.1) from Bacteroides thetaiotaomicron VPI-5482 at 2.40 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.40 Å
R/Rfree: 0.15/0.21
Resolution: 2.40 Å
R/Rfree: 0.15/0.21
X-ray diffraction data for the CRYSTAL STRUCTURE OF A PUTATIVE SUGAR ISOMERASE (LMO2234) FROM LISTERIA MONOCYTOGENES AT 1.70 A RESOLUTION


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.70 Å
R/Rfree: 0.19/0.24
Resolution: 1.70 Å
R/Rfree: 0.19/0.24
X-ray diffraction data for the Crystal structure of a N-acetylmuramoyl-L-alanine amidase (BACUNI_02947) from Bacteroides uniformis ATCC 8492 at 1.07 A resolution


First author:
JOINT CENTER FOR STRUCTURAL GENOMICS (JCSG)
Resolution: 1.07 Å
R/Rfree: 0.11/0.14
Resolution: 1.07 Å
R/Rfree: 0.11/0.14
X-ray diffraction data for the Crystal structure of a pheromone cOB1 precursor/lipoprotein, YaeC family (EF2496) from Enterococcus faecalis V583 at 1.90 A resolution


First author:
Joint center for structural genomics (JCSG)
Resolution: 1.90 Å
R/Rfree: 0.19/0.24
Resolution: 1.90 Å
R/Rfree: 0.19/0.24
X-ray diffraction data for the Crystal structure of Methylglyoxal synthase (TM1185) from Thermotoga maritima at 2.06 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.06 Å
R/Rfree: 0.17/0.22
Resolution: 2.06 Å
R/Rfree: 0.17/0.22
X-ray diffraction data for the Crystal structure of 6-phosphogluconolactonase (TM1154) from Thermotoga maritima at 1.70A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.55 Å
R/Rfree: 0.17/0.20
Resolution: 1.55 Å
R/Rfree: 0.17/0.20
X-ray diffraction data for the Crystal structure of a putative acylhydrolase (BACUNI_03406) from Bacteroides uniformis ATCC 8492 at 1.37 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.37 Å
R/Rfree: 0.13/0.17
Resolution: 1.37 Å
R/Rfree: 0.13/0.17
X-ray diffraction data for the Crystal structure of a pyridoxamine 5'-phosphate oxidase related protein (psyc_0186) from psychrobacter arcticus 273-4 at 2.50 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.50 Å
R/Rfree: 0.19/0.23
Resolution: 2.50 Å
R/Rfree: 0.19/0.23
X-ray diffraction data for the Crystal structure of a centrosomal protein 170kDa, transcript variant beta (CEP170) from Homo sapiens at 2.15 A resolution (PSI Community Target, Sundstrom)


First author:
JOINT CENTER FOR STRUCTURAL GENOMICS (JCSG)
Resolution: 2.15 Å
R/Rfree: 0.18/0.20
Resolution: 2.15 Å
R/Rfree: 0.18/0.20
X-ray diffraction data for the Crystal structure of putative hydrolase (2632844) from Bacillus subtilis at 1.96 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.96 Å
R/Rfree: 0.18/0.24
Resolution: 1.96 Å
R/Rfree: 0.18/0.24
X-ray diffraction data for the Crystal structure of Putative reductase (NP_038806.2) from MUS MUSCULUS at 1.18 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.18 Å
R/Rfree: 0.13/0.16
Resolution: 1.18 Å
R/Rfree: 0.13/0.16
X-ray diffraction data for the Crystal structure of an acyl-ACP thioesterase (NP_810988.1) from Bacteroides thetaiotaomicron VPI-5482 at 1.90 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.90 Å
R/Rfree: 0.17/0.21
Resolution: 1.90 Å
R/Rfree: 0.17/0.21
X-ray diffraction data for the CRYSTAL STRUCTURE OF A PUTATIVE CYSTATHIONINE BETA-LYASE INVOLVED IN ALUMINUM RESISTANCE (LMOF2365_1314) FROM LISTERIA MONOCYTOGENES STR. 4B F2365 AT 1.91 A RESOLUTION


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.91 Å
R/Rfree: 0.21/0.24
Resolution: 1.91 Å
R/Rfree: 0.21/0.24
X-ray diffraction data for the Crystal structure of CT0912, ORFan protein from Chlorobium tepidum with a ferredoxin-like domain repeat (NP_661805.1) from CHLOROBIUM TEPIDUM TLS at 1.80 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.80 Å
R/Rfree: 0.16/0.19
Resolution: 1.80 Å
R/Rfree: 0.16/0.19
X-ray diffraction data for the Crystal structure of Inosine-5'-monophosphate dehydrogenase (TM1347) from THERMOTOGA MARITIMA at 2.18 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.18 Å
R/Rfree: 0.22/0.26
Resolution: 2.18 Å
R/Rfree: 0.22/0.26
X-ray diffraction data for the CRYSTAL STRUCTURE OF a putative nucleic acid binding protein (TM0693) FROM THERMOTOGA MARITIMA MSB8 AT 2.05 A RESOLUTION


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.05 Å
R/Rfree: 0.19/0.25
Resolution: 2.05 Å
R/Rfree: 0.19/0.25
X-ray diffraction data for the Crystal structure of a protein member of the upf0052 family (bh3568) from bacillus halodurans at 2.60 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.60 Å
R/Rfree: 0.19/0.22
Resolution: 2.60 Å
R/Rfree: 0.19/0.22
X-ray diffraction data for the Crystal structure of a putative dihydrofolate reductase (bsu40760, yyap) from bacillus subtilis at 2.30 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.30 Å
R/Rfree: 0.21/0.25
Resolution: 2.30 Å
R/Rfree: 0.21/0.25
X-ray diffraction data for the Crystal structure of an ob-fold protein (tm0957) from thermotoga maritima msb8 at 1.90 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.25 Å
R/Rfree: 0.19/0.25
Resolution: 2.25 Å
R/Rfree: 0.19/0.25
X-ray diffraction data for the Crystal structure of a putative cell adhesion protein (BACOVA_04078) from Bacteroides ovatus ATCC 8483 at 2.80 A resolution


First author:
JOINT CENTER FOR STRUCTURAL GENOMICS (JCSG)
Resolution: 2.80 Å
R/Rfree: 0.20/0.25
Resolution: 2.80 Å
R/Rfree: 0.20/0.25
X-ray diffraction data for the Crystal structure of a predicted dna-binding transcriptional regulator (saro_1072) from novosphingobium aromaticivorans dsm at 2.10 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.10 Å
R/Rfree: 0.17/0.20
Resolution: 2.10 Å
R/Rfree: 0.17/0.20
X-ray diffraction data for the CRYSTAL STRUCTURE OF A TWO-DOMAIN PROTEIN CONTAINING PREDICTED PHP-LIKE METAL-DEPENDENT PHOSPHOESTERASE (BVU_3505) FROM BACTEROIDES VULGATUS ATCC 8482 AT 2.20 A RESOLUTION


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.20 Å
R/Rfree: 0.17/0.23
Resolution: 2.20 Å
R/Rfree: 0.17/0.23
X-ray diffraction data for the Crystal structure of Anthranilate phosphoribosyltransferase 2 (17130499) from Nostoc sp. at 1.85 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.85 Å
R/Rfree: 0.18/0.22
Resolution: 1.85 Å
R/Rfree: 0.18/0.22
X-ray diffraction data for the Crystal structure of a putative phosphomethylpyrimidine kinase (BT_4458) from BACTEROIDES THETAIOTAOMICRON VPI-5482 at 2.00 A resolution (orthorhombic form with pyridoxal)


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.00 Å
R/Rfree: 0.17/0.21
Resolution: 2.00 Å
R/Rfree: 0.17/0.21
X-ray diffraction data for the Crystal structure of an ethyl tert-butyl ether d (ethd) family protein (bh0200) from bacillus halodurans c-125 at 1.40 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.40 Å
R/Rfree: 0.15/0.17
Resolution: 1.40 Å
R/Rfree: 0.15/0.17
X-ray diffraction data for the Crystal structure of a protein of unknown function with a cystatin-like fold (npun_r3134) from nostoc punctiforme pcc 73102 at 1.80 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.80 Å
R/Rfree: 0.16/0.20
Resolution: 1.80 Å
R/Rfree: 0.16/0.20
X-ray diffraction data for the Crystal structure of putative flavin reductase with split barrel domain (YP_750721.1) from SHEWANELLA FRIGIDIMARINA NCIMB 400 at 1.74 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.74 Å
R/Rfree: 0.17/0.21
Resolution: 1.74 Å
R/Rfree: 0.17/0.21
X-ray diffraction data for the Crystal structure of a DUF4467 family protein (SAV0303) from Staphylococcus aureus subsp. aureus Mu50 at 1.35 A resolution

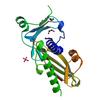
First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.20 Å
R/Rfree: 0.14/0.17
Resolution: 1.20 Å
R/Rfree: 0.14/0.17
X-ray diffraction data for the Crystal structure of acetyltransferase (NP_689019.1) from Streptococcus agalactiae 2603 at 2.00 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.00 Å
R/Rfree: 0.19/0.22
Resolution: 2.00 Å
R/Rfree: 0.19/0.22
X-ray diffraction data for the Crystal structure of a leucine-rich repeat protein (BACCAP_00569) from Bacteroides capillosus ATCC 29799 at 2.00 A resolution


First author:
JOINT CENTER FOR STRUCTURAL GENOMICS (JCSG)
Resolution: 2.00 Å
R/Rfree: 0.17/0.19
Resolution: 2.00 Å
R/Rfree: 0.17/0.19
X-ray diffraction data for the CRYSTAL STRUCTURE OF a DUF574 family protein (SAV_2177) FROM STREPTOMYCES AVERMITILIS MA-4680 AT 1.45 A RESOLUTION


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.45 Å
R/Rfree: 0.16/0.18
Resolution: 1.45 Å
R/Rfree: 0.16/0.18
X-ray diffraction data for the Crystal structure of a putative lipoprotein (BF2707) from Bacteroides fragilis NCTC 9343 at 2.75 A resolution


First author:
JOINT CENTER FOR STRUCTURAL GENOMICS (JCSG)
Resolution: 2.48 Å
R/Rfree: 0.20/0.23
Resolution: 2.48 Å
R/Rfree: 0.20/0.23
X-ray diffraction data for the Crystal structure of a DUF4450 family protein (BT_4147) from Bacteroides thetaiotaomicron VPI-5482 at 2.90 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.90 Å
R/Rfree: 0.19/0.22
Resolution: 2.90 Å
R/Rfree: 0.19/0.22
X-ray diffraction data for the Crystal structure of a BLIP-like protein (BF1215) from Bacteroides fragilis NCTC 9343 at 1.20 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.20 Å
R/Rfree: 0.15/0.17
Resolution: 1.20 Å
R/Rfree: 0.15/0.17
X-ray diffraction data for the Crystal structure of a DUF487 family protein (DESPIG_00776) from Desulfovibrio piger ATCC 29098 at 2.49 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.40 Å
R/Rfree: 0.18/0.23
Resolution: 2.40 Å
R/Rfree: 0.18/0.23
X-ray diffraction data for the Crystal structure of a putative oxidoreductase (YP_511008.1) from Jannaschia sp. CCS1 at 1.62 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.62 Å
R/Rfree: 0.16/0.20
Resolution: 1.62 Å
R/Rfree: 0.16/0.20
X-ray diffraction data for the Crystal structure of Putative PhoU-like phosphate regulatory protein (BT4638) from BACTEROIDES THETAIOTAOMICRON VPI-5482 at 1.93 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.93 Å
R/Rfree: 0.17/0.20
Resolution: 1.93 Å
R/Rfree: 0.17/0.20
X-ray diffraction data for the Crystal structure of a susd homolog (BVU_2203) from Bacteroides vulgatus ATCC 8482 at 1.85 A resolution


First author:
JOINT CENTER FOR STRUCTURAL GENOMICS (JCSG)
Resolution: 1.85 Å
R/Rfree: 0.14/0.16
Resolution: 1.85 Å
R/Rfree: 0.14/0.16
X-ray diffraction data for the Crystal structure of a putative fe(iii) abc transporter (tm0189) from thermotoga maritima msb8 at 1.70 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.70 Å
R/Rfree: 0.16/0.21
Resolution: 1.70 Å
R/Rfree: 0.16/0.21
X-ray diffraction data for the Crystal structure of a protein with unknown function (arth_0117) from arthrobacter sp. fb24 at 2.20 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.20 Å
R/Rfree: 0.18/0.25
Resolution: 2.20 Å
R/Rfree: 0.18/0.25
X-ray diffraction data for the Crystal structure of putative nitroreductase ydfN (2632848) from Bacillus subtilis at 1.65 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.65 Å
R/Rfree: 0.15/0.18
Resolution: 1.65 Å
R/Rfree: 0.15/0.18

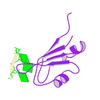
First author:
Partnership for T-Cell Biology (TCELL) Joint Center for Structural Genomics (JCSG)
Resolution: 2.50 Å
R/Rfree: 0.19/0.25
Resolution: 2.50 Å
R/Rfree: 0.19/0.25
X-ray diffraction data for the CRYSTAL STRUCTURE OF A MONOOXYGENASE-LIKE PROTEIN (LIN2316) FROM LISTERIA INNOCUA AT 1.85 A RESOLUTION


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.85 Å
R/Rfree: 0.17/0.19
Resolution: 1.85 Å
R/Rfree: 0.17/0.19
X-ray diffraction data for the Crystal structure of a metallo-endopeptidases (BACOVA_00663) from Bacteroides ovatus at 1.93 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.93 Å
R/Rfree: 0.17/0.21
Resolution: 1.93 Å
R/Rfree: 0.17/0.21
X-ray diffraction data for the CRYSTAL STRUCTURE OF A NTF2-LIKE PROTEIN (AVA_2261) FROM ANABAENA VARIABILIS ATCC 29413 AT 1.65 A RESOLUTION


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.65 Å
R/Rfree: 0.18/0.22
Resolution: 1.65 Å
R/Rfree: 0.18/0.22
X-ray diffraction data for the Crystal structure of an inhibitor of vertebrate lysozyme (PA3902) from Pseudomonas aeruginosa PAO1 at 1.25 A resolution


First author:
JOINT CENTER FOR STRUCTURAL GENOMICS (JCSG)
Resolution: 1.25 Å
R/Rfree: 0.12/0.16
Resolution: 1.25 Å
R/Rfree: 0.12/0.16
X-ray diffraction data for the Crystal structure of a putative acetamidase (tm0119) from thermotoga maritima msb8 at 2.50 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.50 Å
R/Rfree: 0.19/0.24
Resolution: 2.50 Å
R/Rfree: 0.19/0.24
X-ray diffraction data for the Crystal structure of Lmo0035 protein (46906266) from LISTERIA MONOCYTOGENES 4b F2365 at 1.50 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.50 Å
R/Rfree: 0.14/0.17
Resolution: 1.50 Å
R/Rfree: 0.14/0.17
X-ray diffraction data for the Crystal structure of a putative esterase (BDI_1566) from Parabacteroides distasonis ATCC 8503 at 1.60 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.60 Å
R/Rfree: 0.16/0.19
Resolution: 1.60 Å
R/Rfree: 0.16/0.19
X-ray diffraction data for the CRYSTAL STRUCTURE OF A CBS DOMAIN PAIR/ACT DOMAIN PROTEIN (TM0892) FROM THERMOTOGA MARITIMA AT 1.70 A RESOLUTION


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.70 Å
R/Rfree: 0.25/0.27
Resolution: 1.70 Å
R/Rfree: 0.25/0.27
X-ray diffraction data for the Crystal structure of Glucokinase (BDI_1628) from Parabacteroides distasonis ATCC 8503 at 3.00 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 3.00 Å
R/Rfree: 0.21/0.25
Resolution: 3.00 Å
R/Rfree: 0.21/0.25
X-ray diffraction data for the Crystal structure of a FHA domain of kanadaptin (SLC4A1AP) from Homo sapiens at 1.55 A resolution


First author:
JOINT CENTER FOR STRUCTURAL GENOMICS (JCSG)
Resolution: 1.55 Å
R/Rfree: 0.18/0.20
Resolution: 1.55 Å
R/Rfree: 0.18/0.20
X-ray diffraction data for the Crystal structure of Ornithine carbamoyltransferase (TM1097) from Thermotoga maritima at 2.25 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.25 Å
R/Rfree: 0.19/0.22
Resolution: 2.25 Å
R/Rfree: 0.19/0.22
X-ray diffraction data for the Crystal structure of a duf692 family protein (hs_1138) from haemophilus somnus 129pt at 2.20 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.20 Å
R/Rfree: 0.20/0.26
Resolution: 2.20 Å
R/Rfree: 0.20/0.26
X-ray diffraction data for the Crystal structure of a DUF1672 family protein (SAV1486) from Staphylococcus aureus subsp. aureus Mu50 at 1.80 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.80 Å
R/Rfree: 0.18/0.22
Resolution: 1.80 Å
R/Rfree: 0.18/0.22
X-ray diffraction data for the CRYSTAL STRUCTURE OF A NTF2-LIKE PROTEIN (BXE_B1094) FROM BURKHOLDERIA XENOVORANS LB400 AT 1.59 A RESOLUTION


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.59 Å
R/Rfree: 0.18/0.21
Resolution: 1.59 Å
R/Rfree: 0.18/0.21
X-ray diffraction data for the Crystal structure of putative ribokinase (10640157) from Thermoplasma acidophilum at 1.91 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.91 Å
R/Rfree: 0.20/0.25
Resolution: 1.91 Å
R/Rfree: 0.20/0.25
X-ray diffraction data for the Crystal structure of an apoptosis-inducing factor, mitochondrion-associated, 1 (AIFM1) from Homo sapiens at 1.88 A resolution


First author:
Partnership for T-Cell Biology (TCELL) Joint Center for Structural Genomics (JCSG)
Resolution: 1.88 Å
R/Rfree: 0.17/0.21
Resolution: 1.88 Å
R/Rfree: 0.17/0.21
X-ray diffraction data for the Crystal structure of aminopeptidase PepT (NP_980509.1) from Bacillus cereus ATCC 10987 at 2.04 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.04 Å
R/Rfree: 0.17/0.21
Resolution: 2.04 Å
R/Rfree: 0.17/0.21
X-ray diffraction data for the CRYSTAL STRUCTURE OF A PROTEIN WITH A CUPIN-LIKE FOLD AND UNKNOWN FUNCTION (BXE_C0505) FROM BURKHOLDERIA XENOVORANS LB400 AT 1.55 A RESOLUTION


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.55 Å
R/Rfree: 0.17/0.19
Resolution: 1.55 Å
R/Rfree: 0.17/0.19
X-ray diffraction data for the Crystal structure of an anti-ECFsigma factor, ChrR (Maqu_0586) from MARINOBACTER AQUAEOLEI VT8 at 1.70 A resolution

First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.70 Å
R/Rfree: 0.15/0.19
Resolution: 1.70 Å
R/Rfree: 0.15/0.19
X-ray diffraction data for the Crystal structure of a putative conserved lipoprotein (NT01CX_1156) from Clostridium novyi NT at 2.70 A resolution


First author:
JOINT CENTER FOR STRUCTURAL GENOMICS (JCSG)
Resolution: 2.70 Å
R/Rfree: 0.20/0.20
Resolution: 2.70 Å
R/Rfree: 0.20/0.20
X-ray diffraction data for the Crystal structure of a DNA methyltransferase 1 associated protein 1 (DMAP1) from Homo sapiens at 1.45 A resolution


First author:
Partnership for T-Cell Biology Joint Center for Structural Genomics (JCSG)
Resolution: 1.45 Å
R/Rfree: 0.20/0.23
Resolution: 1.45 Å
R/Rfree: 0.20/0.23
X-ray diffraction data for the Crystal structure of a putative zinc-binding metallo-peptidase (BACCAC_01431) from Bacteroides caccae ATCC 43185 at 2.10 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.09 Å
R/Rfree: 0.20/0.23
Resolution: 2.09 Å
R/Rfree: 0.20/0.23
X-ray diffraction data for the Crystal structure of a putative hydrolase (BACOVA_04882) from Bacteroides ovatus ATCC 8483 at 2.80 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.80 Å
R/Rfree: 0.19/0.21
Resolution: 2.80 Å
R/Rfree: 0.19/0.21

First author:
Partnership for T-Cell Biology (TCELL) Joint Center for Structural Genomics (JCSG)
Resolution: 1.75 Å
R/Rfree: 0.18/0.25
Resolution: 1.75 Å
R/Rfree: 0.18/0.25
X-ray diffraction data for the Crystal structure of NudC domain-containing protein 2 (13542905) from Mus musculus at 1.95 A resolution

First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.95 Å
R/Rfree: 0.19/0.24
Resolution: 1.95 Å
R/Rfree: 0.19/0.24
X-ray diffraction data for the Crystal structure of a putative had-like phosphatase (mll2559) from mesorhizobium loti at 1.50 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.50 Å
R/Rfree: 0.14/0.17
Resolution: 1.50 Å
R/Rfree: 0.14/0.17
X-ray diffraction data for the Crystal structure of a TENA/THI-4 domain-containing protein (SSO2700) from Sulfolobus solfataricus P2 at 2.34 A resolution


First author:
JOINT CENTER FOR STRUCTURAL GENOMICS (JCSG)
Resolution: 2.34 Å
R/Rfree: 0.17/0.22
Resolution: 2.34 Å
R/Rfree: 0.17/0.22
X-ray diffraction data for the Crystal structure of a duf416 family protein (maqu_0942) from marinobacter aquaeolei vt8 at 2.00 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.00 Å
R/Rfree: 0.17/0.20
Resolution: 2.00 Å
R/Rfree: 0.17/0.20
X-ray diffraction data for the Crystal structure of a putative dipeptidyl-peptidase VI (BT_1314) from BACTEROIDES THETAIOTAOMICRON VPI-5482 at 2.10 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.10 Å
R/Rfree: 0.17/0.20
Resolution: 2.10 Å
R/Rfree: 0.17/0.20
X-ray diffraction data for the Crystal structure of a pyrophosphatase (AF1178) from Archaeoglobus fulgidus DSM 4304 at 1.80 A resolution


First author:
JOINT CENTER FOR STRUCTURAL GENOMICS (JCSG)
Resolution: 1.78 Å
R/Rfree: 0.19/0.24
Resolution: 1.78 Å
R/Rfree: 0.19/0.24
X-ray diffraction data for the Crystal structure of Argininosuccinate synthase (TM1780) from Thermotoga maritima at 1.65 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.65 Å
R/Rfree: 0.18/0.21
Resolution: 1.65 Å
R/Rfree: 0.18/0.21
X-ray diffraction data for the Crystal structure of a DUF1343 family protein (BF0371) from Bacteroides fragilis NCTC 9343 at 1.65 A resolution


First author:
JOINT CENTER FOR STRUCTURAL GENOMICS (JCSG)
Resolution: 1.65 Å
R/Rfree: 0.15/0.18
Resolution: 1.65 Å
R/Rfree: 0.15/0.18
X-ray diffraction data for the Crystal structure of a PDZ domain protein (BDI_1242) from Parabacteroides distasonis ATCC 8503 at 2.50 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.50 Å
R/Rfree: 0.19/0.22
Resolution: 2.50 Å
R/Rfree: 0.19/0.22
X-ray diffraction data for the Crystal structure of Putative protein binding protein (NP_241345.1) from Bacillus halodurans at 2.71 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.71 Å
R/Rfree: 0.22/0.23
Resolution: 2.71 Å
R/Rfree: 0.22/0.23
X-ray diffraction data for the CRYSTAL STRUCTURE OF A PUTATIVE SERINE HYDROLASE (XCC3885) FROM XANTHOMONAS CAMPESTRIS PV. CAMPESTRIS AT 2.69 A RESOLUTION

First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.69 Å
R/Rfree: 0.20/0.22
Resolution: 2.69 Å
R/Rfree: 0.20/0.22
X-ray diffraction data for the Crystal structure of uncharacterized protein (YP_323524.1) from Anabaena variabilis ATCC 29413 at 2.11 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.11 Å
R/Rfree: 0.19/0.22
Resolution: 2.11 Å
R/Rfree: 0.19/0.22
X-ray diffraction data for the Crystal structure of glycerol dehydrogenase (NP_348253.1) from Clostridium acetobutylicum at 2.37 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.37 Å
R/Rfree: 0.19/0.21
Resolution: 2.37 Å
R/Rfree: 0.19/0.21
X-ray diffraction data for the Crystal structure of Putative phosphatase (DUF442) (YP_001181608.1) from SHEWANELLA PUTREFACIENS CN-32 at 1.40 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.40 Å
R/Rfree: 0.16/0.18
Resolution: 1.40 Å
R/Rfree: 0.16/0.18
X-ray diffraction data for the Crystal structure of a putative 6-phosphogluconolactonase (BF1038) from Bacteroides fragilis NCTC 9343 at 1.50 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.50 Å
R/Rfree: 0.16/0.19
Resolution: 1.50 Å
R/Rfree: 0.16/0.19
X-ray diffraction data for the Crystal structure of a putative carboxylesterase (lp_1002) from lactobacillus plantarum wcfs1 at 2.09 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.09 Å
R/Rfree: 0.18/0.19
Resolution: 2.09 Å
R/Rfree: 0.18/0.19
X-ray diffraction data for the Crystal structure of a bacterial lipoprotein (bt_1233) from bacteroides thetaiotaomicron vpi-5482 at 1.69 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.69 Å
R/Rfree: 0.16/0.21
Resolution: 1.69 Å
R/Rfree: 0.16/0.21
X-ray diffraction data for the Crystal structure of a cystatin-like protein (saro_2766) from novosphingobium aromaticivorans dsm at 1.40 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.40 Å
R/Rfree: 0.14/0.16
Resolution: 1.40 Å
R/Rfree: 0.14/0.16
X-ray diffraction data for the Crystal structure of an internalin C2 (inlC2) from Listeria monocytogenes str. 4b F2365 at 1.90 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.90 Å
R/Rfree: 0.19/0.23
Resolution: 1.90 Å
R/Rfree: 0.19/0.23
X-ray diffraction data for the Crystal structure of a sugar ABC transporter (ACTODO_00688) from Actinomyces odontolyticus ATCC 17982 at 2.60 A resolution


First author:
JOINT CENTER FOR STRUCTURAL GENOMICS (JCSG)
Resolution: 2.60 Å
R/Rfree: 0.21/0.22
Resolution: 2.60 Å
R/Rfree: 0.21/0.22